Filter News
Area of Research
- (-) National Security (14)
- (-) Neutron Science (11)
- (-) Supercomputing (31)
- Advanced Manufacturing (1)
- Biological Systems (1)
- Biology and Environment (18)
- Clean Energy (36)
- Computational Biology (2)
- Computational Engineering (1)
- Computer Science (7)
- Electricity and Smart Grid (2)
- Fusion and Fission (4)
- Fusion Energy (6)
- Isotopes (4)
- Materials (11)
- Materials for Computing (3)
- Nuclear Science and Technology (12)
- Nuclear Systems Modeling, Simulation and Validation (1)
- Quantum information Science (7)
- Sensors and Controls (1)
News Type
News Topics
- (-) Advanced Reactors (1)
- (-) Biomedical (16)
- (-) Grid (6)
- (-) Machine Learning (16)
- (-) Quantum Science (15)
- 3-D Printing/Advanced Manufacturing (5)
- Artificial Intelligence (28)
- Big Data (20)
- Bioenergy (7)
- Biology (9)
- Biotechnology (2)
- Buildings (2)
- Chemical Sciences (3)
- Clean Water (2)
- Climate Change (17)
- Computer Science (68)
- Coronavirus (11)
- Critical Materials (3)
- Cybersecurity (9)
- Decarbonization (5)
- Energy Storage (7)
- Environment (24)
- Exascale Computing (13)
- Fossil Energy (1)
- Frontier (14)
- Fusion (1)
- High-Performance Computing (25)
- Materials (14)
- Materials Science (17)
- Mathematics (1)
- Microscopy (3)
- Nanotechnology (8)
- National Security (23)
- Net Zero (1)
- Neutron Science (58)
- Nuclear Energy (6)
- Physics (5)
- Polymers (3)
- Quantum Computing (14)
- Security (7)
- Simulation (11)
- Software (1)
- Space Exploration (4)
- Summit (27)
- Sustainable Energy (5)
- Transportation (7)
Media Contacts
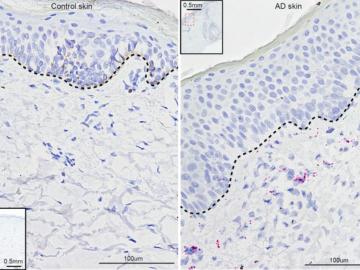
University of Pennsylvania researchers called on computational systems biology expertise at Oak Ridge National Laboratory to analyze large datasets of single-cell RNA sequencing from skin samples afflicted with atopic dermatitis.

A rapidly emerging consensus in the scientific community predicts the future will be defined by humanity’s ability to exploit the laws of quantum mechanics.
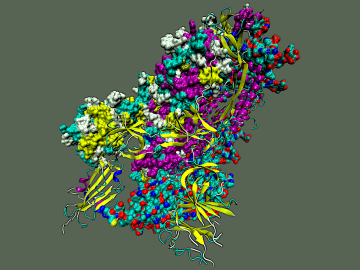
To explore the inner workings of severe acute respiratory syndrome coronavirus 2, or SARS-CoV-2, researchers from ORNL developed a novel technique.
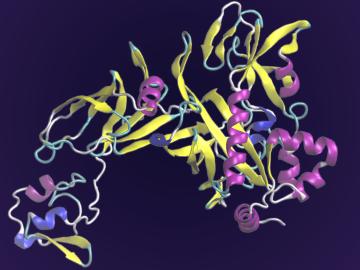
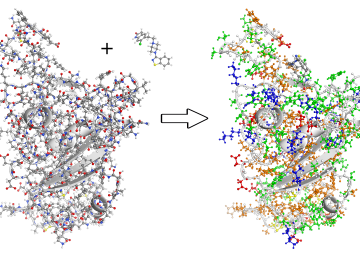
A team of scientists led by the Department of Energy’s Oak Ridge National Laboratory and the Georgia Institute of Technology is using supercomputing and revolutionary deep learning tools to predict the structures and roles of thousands of proteins with unknown functions.
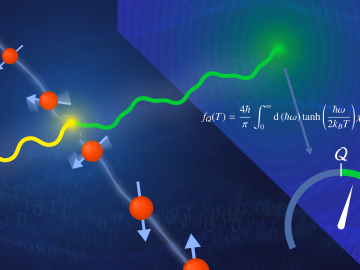
A team led by the U.S. Department of Energy’s Oak Ridge National Laboratory demonstrated the viability of a “quantum entanglement witness” capable of proving the presence of entanglement between magnetic particles, or spins, in a quantum material.
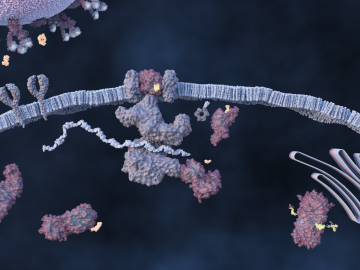
An ORNL-led team comprising researchers from multiple DOE national laboratories is using artificial intelligence and computational screening techniques – in combination with experimental validation – to identify and design five promising drug therapy approaches to target the SARS-CoV-2 virus.

Deborah Frincke, one of the nation’s preeminent computer scientists and cybersecurity experts, serves as associate laboratory director of ORNL’s National Security Science Directorate. Credit: Carlos Jones/ORNL, U.S. Dept. of Energy
To better understand the spread of SARS-CoV-2, the virus that causes COVID-19, Oak Ridge National Laboratory researchers have harnessed the power of supercomputers to accurately model the spike protein that binds the novel coronavirus to a human cell receptor.

A multi-institutional team, led by a group of investigators at Oak Ridge National Laboratory, has been studying various SARS-CoV-2 protein targets, including the virus’s main protease. The feat has earned the team a finalist nomination for the Association of Computing Machinery, or ACM, Gordon Bell Special Prize for High Performance Computing-Based COVID-19 Research.
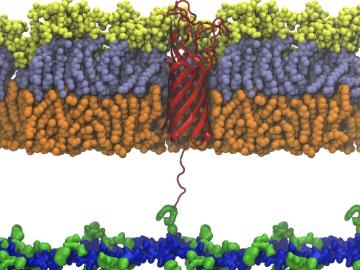
Scientists from Oak Ridge National Laboratory used high-performance computing to create protein models that helped reveal how the outer membrane is tethered to the cell membrane in certain bacteria.




