Filter News
Area of Research
News Topics
- (-) Artificial Intelligence (29)
- (-) Isotopes (17)
- 3-D Printing/Advanced Manufacturing (44)
- Advanced Reactors (10)
- Big Data (8)
- Bioenergy (24)
- Biology (22)
- Biomedical (17)
- Biotechnology (7)
- Buildings (13)
- Chemical Sciences (29)
- Clean Water (1)
- Climate Change (22)
- Composites (9)
- Computer Science (57)
- Coronavirus (17)
- Critical Materials (11)
- Cybersecurity (17)
- Decarbonization (18)
- Education (3)
- Element Discovery (1)
- Energy Storage (41)
- Environment (36)
- Exascale Computing (9)
- Fossil Energy (1)
- Frontier (14)
- Fusion (14)
- Grid (15)
- High-Performance Computing (26)
- ITER (2)
- Machine Learning (13)
- Materials (59)
- Materials Science (50)
- Mercury (2)
- Microscopy (16)
- Molten Salt (2)
- Nanotechnology (26)
- National Security (18)
- Net Zero (3)
- Neutron Science (49)
- Nuclear Energy (25)
- Partnerships (27)
- Physics (24)
- Polymers (12)
- Quantum Computing (9)
- Quantum Science (26)
- Renewable Energy (1)
- Security (11)
- Simulation (8)
- Space Exploration (3)
- Statistics (2)
- Summit (20)
- Sustainable Energy (31)
- Transformational Challenge Reactor (4)
- Transportation (24)
Media Contacts

OAK RIDGE, Tenn., Jan. 31, 2019—A new electron microscopy technique that detects the subtle changes in the weight of proteins at the nanoscale—while keeping the sample intact—could open a new pathway for deeper, more comprehensive studies of the basic building blocks of life.
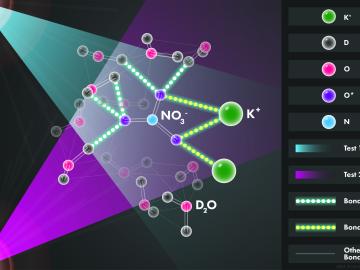
Scientists at the Department of Energy’s Oak Ridge National Laboratory used neutrons, isotopes and simulations to “see” the atomic structure of a saturated solution and found evidence supporting one of two competing hypotheses about how ions come

The Department of Energy’s Oak Ridge National Laboratory is now producing actinium-227 (Ac-227) to meet projected demand for a highly effective cancer drug through a 10-year contract between the U.S. DOE Isotope Program and Bayer.

A team of researchers from the Department of Energy’s Oak Ridge National Laboratory has married artificial intelligence and high-performance computing to achieve a peak speed of 20 petaflops in the generation and training of deep learning networks on the




