A multidisciplinary ORNL team used expertise in synthetic biology, AI-driven analysis, chemistry, neutrons and materials science to identify new members of a family of enzymes with a natural affinity for degrading synthetic nylon polymers.

A multidisciplinary ORNL team used expertise in synthetic biology, AI-driven analysis, chemistry, neutrons and materials science to identify new members of a family of enzymes with a natural affinity for degrading synthetic nylon polymers.

A multidisciplinary team of researchers from Oak Ridge National Laboratory (ORNL) and other institutions created a Machine Learning (ML) library for the training of classifiers on spectrographic chemical data.

Evaluate the historical performance and future projections of compound heatwave and drought (CHD) extremes across the contiguous United States using CMIP6 global climate models, providing insights for regional adaptation strategies in response to

The objective of this study is to explore and analyze the spatial patterning of sociodemographic disparities in extreme heat exposure across multiple scales within the Conterminous United States (CONUS).

A multidisciplinary team of researchers from Oak Ridge National Laboratory (ORNL) pioneered the use of the LLVM-based high-productivity/high-performance Julia language unifying capabilities to write an end-to-end workflow on Frontier, the first US Depar

Computational scientists and neutron structural biologists from Oak Ridge National Laboratory developed an integrated workflow using small-angle neutron scattering (SANS), atomistic molecular dynamics (MD) simulation, and an autoencoder-based deep learn

Large amounts of longitudinal, multimodal electronic health data are being produced from a variety of sources daily. If leveraged properly, these comprehensive data sources can be used for innovative precision medicine and precision public health.

A team of researchers from Oak Ridge National Laboratory (ORNL) released the initial draft of the Interconnected Science Ecosystem (INTERSECT) architecture specification.
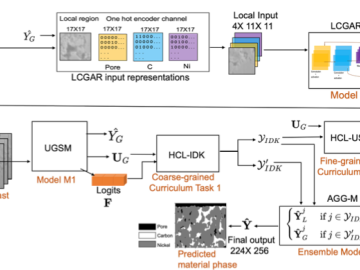
We developed a novel uncertainty-aware framework MatPhase to predict material phases of electrodes from low contrast SEM images.
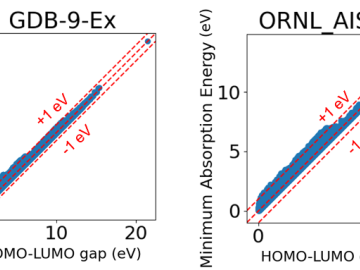
We released two open-source datasets named GDB-9-Ex and ORNL_AISD-Ex that provide calculations of electronic excitation energies and their associated oscillator strengths based on the time-dependent density-functional tight-binding (TD-DFTB) method.

