Achievement: Devised a novel and accurate computational technique for investigating the self-assembly of large macromolecules, and used this method to reveal the initial stages of self-assembly of the carboxysome, the prototype bacterial micro-compartment.
Significance and Impact: Understanding the self-assembly of bacterial micro-compartments can help designing artificial nano-reactors and membranes.
Research Details:
- Atomistic interactions between two CcmK2 hexamers, a major structural component of a carboxysome’s facet, were used to developed a coarse grain (CG) model.
- The CG model was subsequently employed to perform Monte Carlo simulations of CcmK2 self-assembly in 2D.
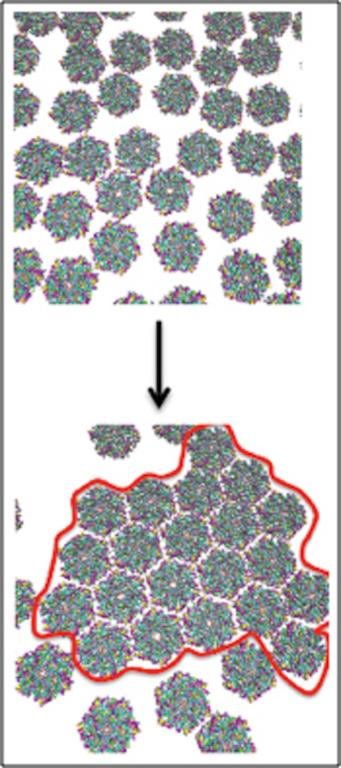
- The simulations revealed that CcmK2 hexamers self-assemble into clusters that resemble those seen experimentally.
- It is concluded that the self-assembly of a carboxysome’s facet is driven by a nucleation-growth process.
Sponsor/Facility: This research was conducted at the Center for Nanophase Materials Sciences (CNMS), which is a U.S. Department of Energy Office of Science User Facility. K. Brown was supported by an appointment under the Science Undergraduate Laboratory Internships (SULI), administered by the Oak Ridge Institute for Science and Education between the U.S. Department of Energy and Oak Ridge Associated Universities. The computations were performed using resources of the CNMS and the Oak Ridge Leadership Computing Facility at Oak Ridge National Laboratory. The authors would like to thank Mitch Doktycz, David Garcia, Pat Collier, Scott Retterer, Rajeev Kumar and Bobby Sumpter for insightful discussions.
PI and affiliation: J. P. Mahalik from CSMD/CNMS – Oak Ridge National Laboratory
Team: J. P. Mahalik, Kirsten A. Brown, Xiaolin Cheng, and Miguel Fuentes-Cabrera
Overview:
Bacterial microcompartments, BMCs, are organelles that exist within certain type of bacteria and act as nano-factories. Among the different types of known BMCs, the carboxysome has been studied the most. The carboxysome plays an important role in the light-independent part of the photosynthesis process, where its icosahedral-like proteinaceous shell acts as a membrane that controls the transport of metabolites. Although a structural model exists for the carboxysome shell, it remains largely unknown how the shell proteins self-assemble. Understanding the self-assembly process can provide insights into how the shell affects the carboxysome's function and how it can be modified to create new functionalities, such as artificial nanoreactors and artificial protein membranes. Here, we describe a theoretical framework that employs Monte Carlo simulations with a coarse grain potential that reproduces well the atomistic potential of mean force; employing this framework, we are able to capture the initial stages of the 2D self-assembly of CcmK2 hexamers, a major protein-shell component of the carboxysome's facet.
The simulations reveal that CcmK2 hexamers self-assemble into clusters that resemble what was seen experimentally in 2D layers. Further analysis of the simulation results suggests that the 2D self-assembly of carboxysome's facets is driven by a nucleation-growth process, which in turn could play an important role in the hierarchical self-assembly of BMCs shells in general.

