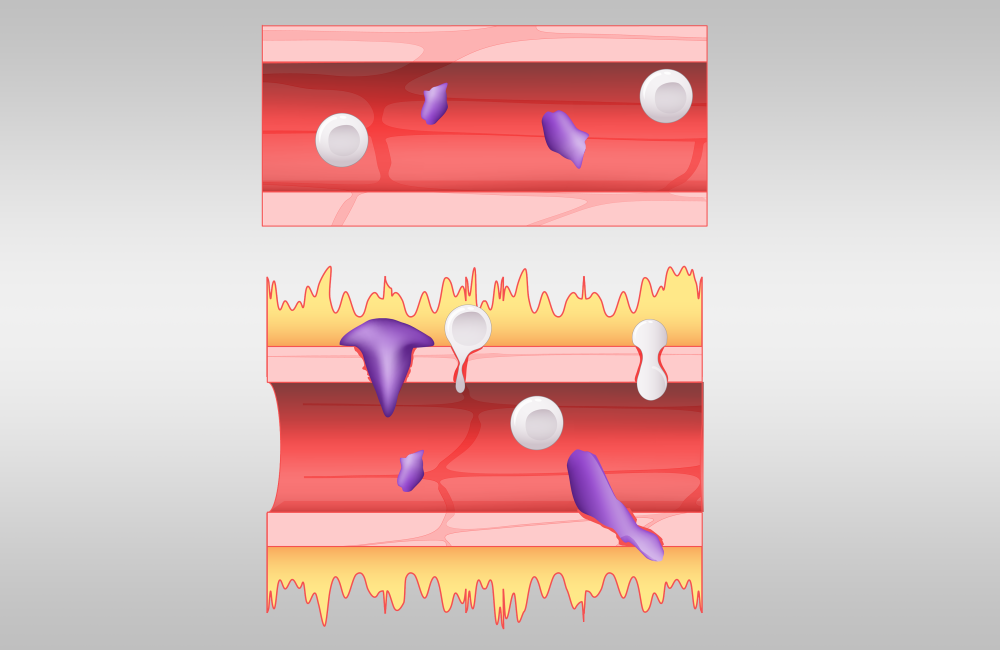
A normal blood vessel, shown at top, is compared with a blood vessel affected by excess bradykinin. A hyperactive bradykinin system permits fluid, shown in yellow, to leak out and allows immune cells, shown in purple, to squeeze their way out of blood vessels. Credit: Jason Smith/ORNL, U.S. Dept. of Energy
Analyses of lung fluid cells from COVID-19 patients conducted on the nation’s fastest supercomputer point to gene expression patterns that may explain the runaway symptoms produced by the body’s response to SARS-CoV-2.
A team led by Dan Jacobson of the Department of Energy’s Oak Ridge National Laboratory used the Summit supercomputer at ORNL to analyze genes from cells in the lung fluid of nine COVID-19 patients compared with 40 control patients.
The computational analyses suggest that genes related to one of the body’s systems responsible for lowering blood pressure — the bradykinin system — appear to be excessively “turned on” in the lung fluid cells of those with the virus. The results were published in eLife.
Based on their analyses, the team posits that bradykinin — the compound that dilates blood vessels and makes them permeable — is overproduced in the body of COVID-19 patients; related systems either contribute to overproduction or cannot slow the process. Excessive bradykinin leads to leaky blood vessels, allowing fluid to build up in the body’s soft tissues.
Much attention has focused on what’s known as the cytokine storm, a severe reaction in which the body releases an excess of cytokines, a variety of small proteins that help regulate the immune system. Jacobson’s team thinks a bradykinin storm may instead be to blame for much of the viral pathogenesis. If the team’s disease mechanism model is accurate and substantiated by experimental analysis, it may mean that existing medicines could be repurposed to slow the pathogenesis of COVID-19. This would require extensive clinical trials of drugs currently used to treat other bradykinin-related conditions.
“If we can block this pathogenesis in severe patients, we can keep the human response from going overboard and give their immune system time to fight off the virus so they can recover,” Jacobson said.
The bradykinin storm could explain the wide variety of symptoms experienced by COVID-19 patients, such as muscle pain, fatigue, nausea, vomiting, diarrhea, headaches, and decreased cognitive function. Similar symptoms are also experienced by patients with other bradykinin-related conditions such as hereditary angioedema, a genetic condition that is characterized by episodes of severe swelling throughout the body.
“This is one of those rare times where you can really tie everything back to a eureka moment,” said Jacobson, staff scientist in ORNL’s Biosciences Division. “I was looking at data, and I suddenly saw some very distinct patterns happening in the pathways of the renin-angiotensin and bradykinin systems. That led us to do a deep dive of the gene families of the blood pressure regulatory system.” The renin-angiotensin system, or RAS, and bradykinin pathway regulate blood pressure and fluid balance in the body.
Using the Summit and Rhea supercomputers at the Oak Ridge Leadership Computing Facility, the team compared the genes of COVID-19 patients against a control group and analyzed population-scale gene expression data — 17,000 samples from uninfected individuals — to see which genes were normally co-expressed, or turned on or off at the same time.
Summit is currently the nation’s most powerful supercomputer, with a theoretical peak performance of 200 petaflops, or 200 quadrillion calculations per second. Jacobson and his colleagues required the power of Summit to run 2.5 billion correlation calculations that helped them understand the normal regulatory circuits and relationships for the genes of interest. With Summit, the team completed the calculations in one week rather than spending months doing them on a desktop computer.
Researchers found an increased expression of enzymes that can trigger the production of bradykinin and a decreased expression of enzymes that would break down bradykinin — the perfect storm. The team also uncovered that an enzyme that forestalls the bradykinin cascade — the angiotensin-converting enzyme, known as ACE — was less expressed in COVID-19 patients. At least ten existing drugs are known to act on the specific pathways Jacobson’s team studied, but large-scale clinical trials are needed to determine whether they might be effective at treating COVID-19.
“We believe that when you take the inhibition at the top of this pathway off, you end up with an out-of-control cascade that leads to an opening up of the blood vessels, causing them to leak,” Jacobson said. “If that happens in the lung, that’s not good. Immune cells that are normally contained in the blood vessels flood into the surrounding infected tissue, causing inflammation.”
The lungs of COVID-19 patients are known to have an increased amount of hyaluronic acid, a gooey substance found in connective tissues that can trap around 1,000 times its own weight in water to form a hydrogel. The team also found that genes in the cells of COVID-19 patients increased the production of the substance and decreased its breakdown. The findings suggest that further experimental study of drug compounds known to slow the synthesis of hyaluronic acid and the mechanisms involved in the process is warranted.
“When the lungs end up with an excess of hyaluronic acid in them, it’s like trying to breathe through Jell-O,” Jacobson said. “It reaches a point where regardless of how much oxygen you pump in, it doesn’t matter, because the alveoli in the lungs are filled with this hydrogel. With this excess of hyaluronic acid, any water leaking out of the blood vessels due to bradykinin will soak up this structure and the lungs become like a water balloon.”
The team also used the Compute and Data Environment for Science, or CADES, at ORNL to determine which genes in the RAS-bradykinin pathways have vitamin D binding sites. The results of their analyses might help scientists determine through experimentation which parts of these pathways could potentially be influenced by vitamin D. Because vitamin D helps regulate the RAS and vitamin D deficiencies have already been associated with more severe illness in COVID-19 patients, Jacobson said it's another molecule worth further study.
The research team included colleagues from ORNL, University of Tennessee, Knoxville, the Yale School of Medicine, the U.S. Department of Veterans Affairs, the Versiti Blood Research Institute, the University of Kentucky and the Cincinnati Children’s Hospital Medical Center.
The OLCF is a DOE Office of Science user facility located at ORNL.
The team’s research was supported by ORNL’s Laboratory Directed Research and Development Program; the DOE Office of Science through the National Virtual Biotechnology Laboratory, a consortium of DOE national laboratories focused on response to COVID-19, with funding provided by the Coronavirus CARES Act; and the National Institutes of Health.
Learn more about ORNL research in the fight against COVID-19.
UT-Battelle manages ORNL for the Department of Energy’s Office of Science, the single largest supporter of basic research in the physical sciences in the United States. The Office of Science is working to address some of the most pressing challenges of our time. For more information, please visit energy.gov/science. — Rachel Harken