
Case closed: Neutrons settle 40-year debate on enzyme for drug design
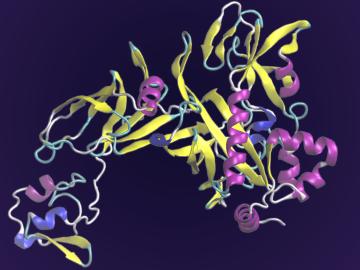
A team of scientists led by the Department of Energy’s Oak Ridge National Laboratory and the Georgia Institute of Technology is using supercomputing and revolutionary deep learning tools to predict the structures and roles of thousands of proteins with unknown functions.
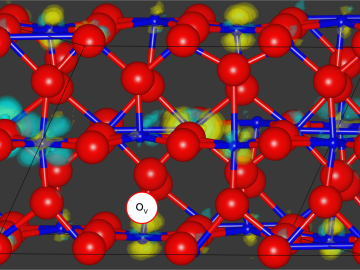
Neuromorphic devices — which emulate the decision-making processes of the human brain — show great promise for solving pressing scientific problems, but building physical systems to realize this potential presents researchers with a significant

