Human Genome Project
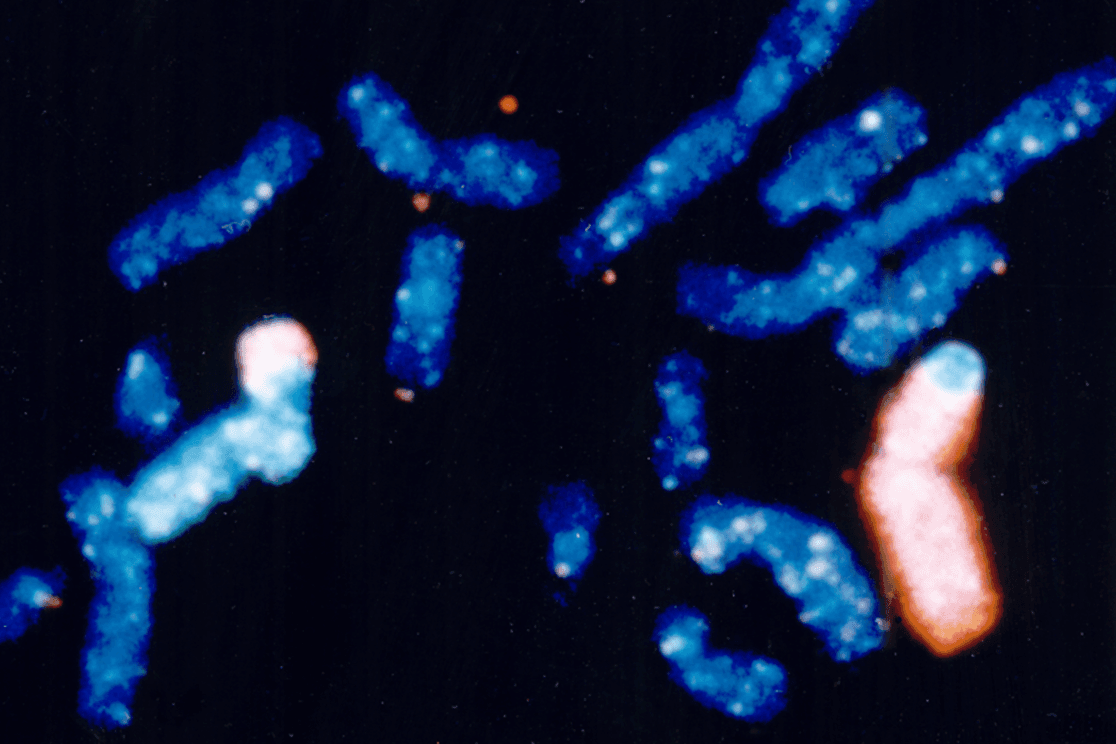
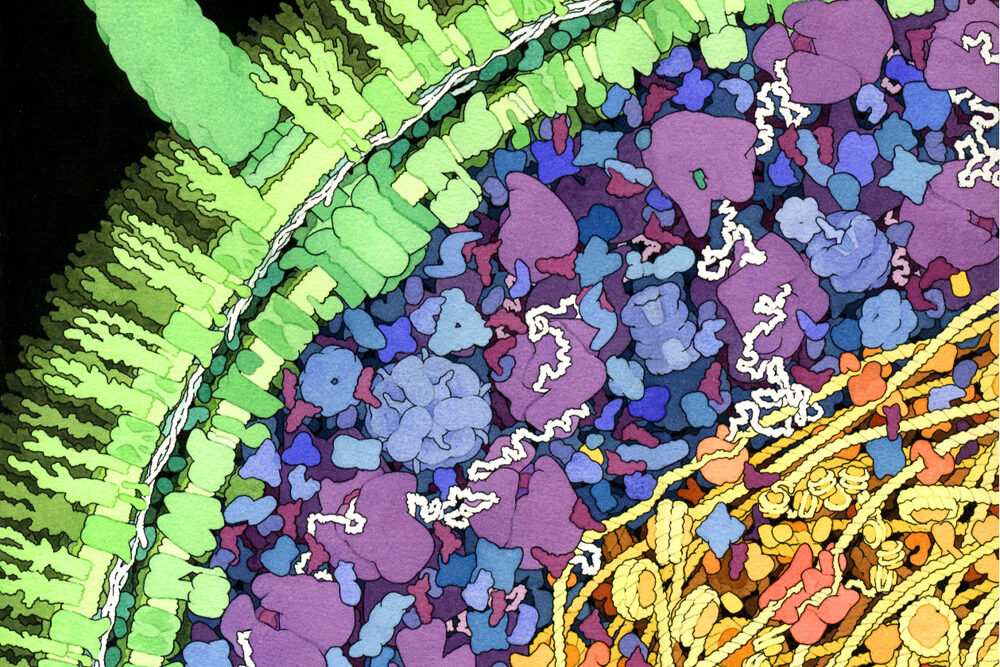
Completed in 2003, the Human Genome Project (HGP) was a 13-year project coordinated by the U.S. Department of Energy (DOE) and the National Institutes of Health. During the early years of the HGP, the Wellcome Trust (U.K.) became a major partner; additional contributions came from Japan, France, Germany, China, and others. This website details HGP history.